RSEM (RNA-Seq by Expectation Maximization) is a software package that quantifies transcript abundances. HISAT2 (Hierarchical Indexing for Spliced Alignment of Transcripts 2) is also a splice-aware aligner using a graph-based alignment approach (graph Ferragina Manzini index) that can align DNA and RNA sequences. STAR (Spliced Transcripts Alignment to a Reference) is a specialized tool for RNA-Seq reads that uses a seed-extension search based on compressed suffix arrays and can detect splice-junctions. For indexing, the algorithm constructs a suffix array and Burrows–Wheeler-Transformation (BWT), and subsequently matches the sequences using a backward search. In this study, the algorithmically different mappers bwa, CLC Genomics Workbench, HISAT2, kallisto, RSEM, salmon and STAR were used to map experimentally generated RNA-Seq data from the two natural accessions Columbia-0 (Col-0) and N14 of the higher plant Arabidopsis thaliana and to quantify the transcripts.īwa (Burrows–Wheeler-Alignment) was developed for mapping short DNA sequences against a reference genome and was extended for RNA-Seq data analysis. While comparative studies of diverse read aligners have been performed using data with a corresponding reference genome or transcriptome or de novo assembly, only little evaluation is available of the performance of read mappers for data generated from genotypes within a species showing sequence polymorphisms. This approach allows detailed transcript profiling including the identification of splicing-induced isoforms, nucleotide variation and post-transcriptional base modification. These two key aspects of transcriptomics can be combined in a single high-throughput sequencing assay called RNA-Sequencing (RNA-Seq). In this context, transcript identification and the quantification of gene expression play crucial roles in connecting genomic information with phenotypic and biochemical measurements. The resulting data have revealed the astonishing complexity of genome architecture and transcriptome composition. Since the completion of the human genome project in 2003, sequencing technologies have developed extraordinarily fast. All tested mappers provided highly similar results for mapping Illumina reads of two polymorphic Arabidopsis accessions to the reference genome or transcriptome and for the determination of DGE when the same software was used for processing. Interestingly, when the commercial CLC software was used with its own DGE module instead of DESeq2, strongly diverging results were obtained. Using the software DESeq2 to determine differential gene expression (DGE) between plants exposed to 20 ☌ or 4 ☌ from these read counts showed a large pairwise overlap between the mappers. Between 92.4% and 99.5% of all reads were mapped to the reference genome or transcriptome and the raw count distributions obtained from the different mappers were highly correlated. Here, we compared seven computational tools for their ability to map and quantify Illumina single-end reads from the Arabidopsis thaliana accessions Columbia-0 (Col-0) and N14.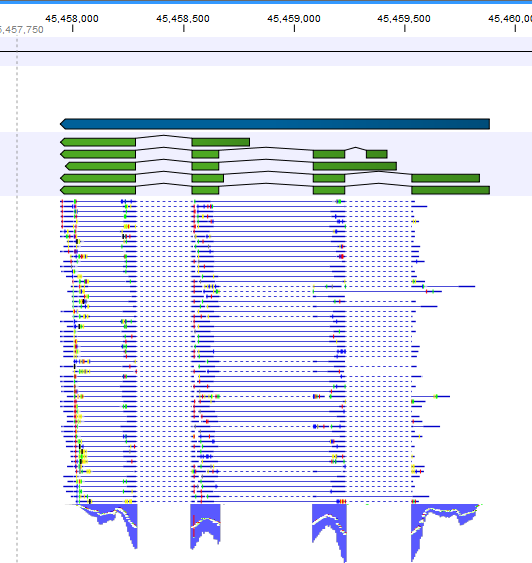 
However, comparative tests of different tools for RNA-Seq read mapping and quantification have been mainly performed on data from animals or humans, which necessarily neglect, for example, the large genetic variability among natural accessions within plant species. RNA-Sequencing (RNA-Seq) has taken a prominent role in the study of transcriptomic reactions of plants to various environmental and genetic perturbations. Quantification of gene expression is crucial to connect genome sequences with phenotypic and physiological data. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
